This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is Proteomics?
Proteomics has become a huge tool in understanding biological functions and their links to genetic diseases, like psoriasis. Proteomics is essentially the study a protein's structure and function. It involves classifying a protein's size, charge, interaction network, function, cellular localization, ect. Understanding these properties provides insight on how a protein functions and interacts with other molecules. Listed below are general protein properties of HLA-C including molecular weight, isoelectric point, and trypsin digest sites.
Molecular Weight (Mw)
http://bio-nin.com/Eng/ACE.htm
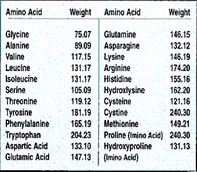
Molecular weight (Mw) is the weight (measured in daltons, Da) of all atoms that make up a protein. Protein Mw is calculated via their amino acid composition. Amino acids are made of of various weights due to their varied atom composition. Mw can be calculated by hand using this table and the protein sequence or can be done using bioinformatic databases, such as ExPASy compute pI/Mw.
HLA-C is comprised of 366 amino acids with an estimated (using the ExPASy) Mw of 40,986.81 daltons (approx. 41 kDa).
HLA-C is comprised of 366 amino acids with an estimated (using the ExPASy) Mw of 40,986.81 daltons (approx. 41 kDa).
Isoelectric Point (pI)
http://www.chemistrypictures.org/index.php/amino-acids/2Electrically_charged
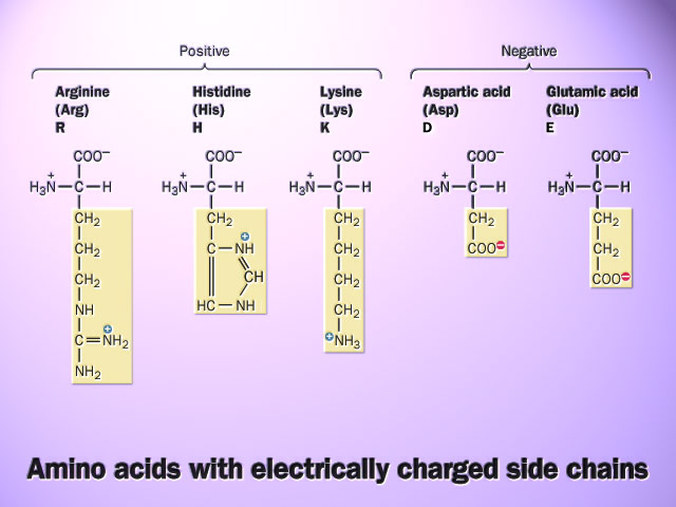
An isoelectric point (pI) is the pH where a molecule has no net charge, meaning that the negative and positive charges balance. pIs values are determined by the amino acids that make up a protein. Amino acids have various charges (see figure) and when combined together in a peptide, the overall charge can be neutral, positive (basic), or negative (acidic). The net charge of a protein or protein fragment affects its solubility, or ability to interact with water. The lowest solubility occurs at its pI, or in other words charges make a protein more soluble because a protein will prefer to interact with water rather than other proteins/molecules. The ExPASy compute pI/Mw can determine the pIs of domains based off their sequences. HLA-C's ExPASy domains are listed below or by clicking here.
Location Domain pI
1-24 Signal 9.35
25-114 Alpha-1 6.8
115-206 Alpha-2 5.09
207-298 Alpha-3 5.18
299-308 Connecting Peptide 4.0
309-333 Transmembrane (Helical) 5.49
334-366 Cytoplasmic (Tail) 9.39
25-366 Overall Chain 5.57
Location Domain pI
1-24 Signal 9.35
25-114 Alpha-1 6.8
115-206 Alpha-2 5.09
207-298 Alpha-3 5.18
299-308 Connecting Peptide 4.0
309-333 Transmembrane (Helical) 5.49
334-366 Cytoplasmic (Tail) 9.39
25-366 Overall Chain 5.57
Trypsin Digests
http://chempaths.chemeddl.org/services/chempaths/
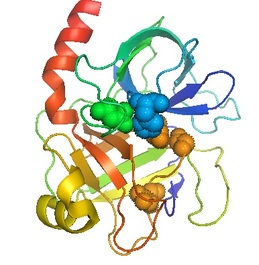
Trypsin (pictured to the left) is a protein that degrades other proteins into smaller pieces. This enzymatic mechanism works by hydrolyzing the carboxyl end of the amino acids, lysine and arginine, causing peptide chains to break apart at specific locations.
In research, trypsin is a biotechnological tool used for proteomic analysis. Proteins are analyzed using a technique call Mass Spectrometry (MS), which determines the compositions of molecules; however MS analyses must analyze proteins in pieces (peptides) because proteins are often too large. Trypsin is used to create these peptide pieces at known locations allowing accurate MS analysis.
ExPASy PeptideCutter is a bioinformatic tool which predicts trypsin digest sites. ExPASy determined 39 cleavage sites for the HLA-C (366 amino acid) protein. The location of the sites can be seen by clicking here.
In research, trypsin is a biotechnological tool used for proteomic analysis. Proteins are analyzed using a technique call Mass Spectrometry (MS), which determines the compositions of molecules; however MS analyses must analyze proteins in pieces (peptides) because proteins are often too large. Trypsin is used to create these peptide pieces at known locations allowing accurate MS analysis.
ExPASy PeptideCutter is a bioinformatic tool which predicts trypsin digest sites. ExPASy determined 39 cleavage sites for the HLA-C (366 amino acid) protein. The location of the sites can be seen by clicking here.
References
Site created by Valeri Lapacek
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
