This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is phylogeny?
Phylogeny shows evolutionary relationships by comparing DNA/protein sequences. Phylogeny is displayed in phylogenetic trees (like a ancestral family tree) which shows which organism (with that DNA/protein sequence) is most closely related.
Protein Phylogeny Analysis
Protein sequences from 5 vertebrate homologs were aligned using various algorithms including: T-coffee, MUSCLE, and ClustalW2. These protein sequence alignments were then used to construct phylogenetic trees using T-coffee, GeneBee, MUSCLE, and ClustalW2. The trees show how the homolog protein sequences, which are conserved to some degree with the human HLA-C protein sequence, are related.
The human and chimpanzee have the most closely related protein sequence, followed by the rat and mouse, being the next related protein sequences. The chicken and the zebrafish have the least conservation among.
All 6 protein phylogeny trees were identical, omitting the distances. Comparing the protein phylogeny to the gene phylogeny, the trees are very similar with the human and chimpanzee being the most closely related, and they zebrafish being the most distant homolog.
The human and chimpanzee have the most closely related protein sequence, followed by the rat and mouse, being the next related protein sequences. The chicken and the zebrafish have the least conservation among.
All 6 protein phylogeny trees were identical, omitting the distances. Comparing the protein phylogeny to the gene phylogeny, the trees are very similar with the human and chimpanzee being the most closely related, and they zebrafish being the most distant homolog.
Phylogeny Trees
GeneBee (ClustalW2 alignment)
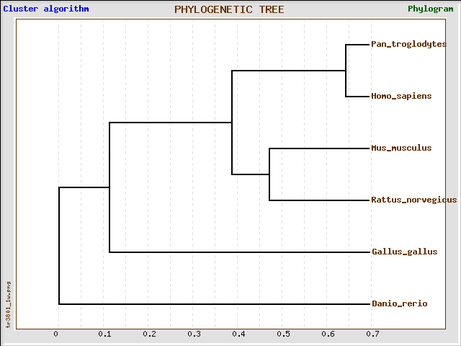
Phylogeny tree constructed with GeneBee.
Sequence alignment data (using ClustalW2) can be visualized here.
Parameters: Pairwise Alignment Options: Protein Weight Matrix: BLOSUM
Multiple Sequence Alignment Options: Protein Weight Matrix: BLOSUM
Sequence alignment data (using ClustalW2) can be visualized here.
Parameters: Pairwise Alignment Options: Protein Weight Matrix: BLOSUM
Multiple Sequence Alignment Options: Protein Weight Matrix: BLOSUM
GeneBee (T-coffee alignment)
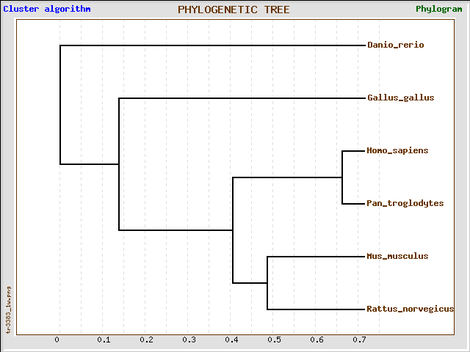
Phylogeny tree constructed with GeneBee.
Sequence alignment data was conducted using T-Coffee. This data can be visualized here.
Sequence alignment data was conducted using T-Coffee. This data can be visualized here.
GeneBee (MUSCLE alignment)
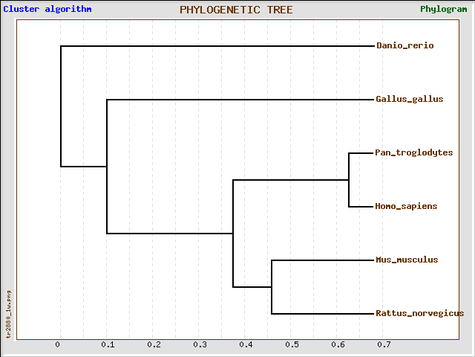
Phylogeny tree constructed with GeneBee.
Sequence alignment data was conducted using MUSCLE. This data can be visualized here.
Sequence alignment data was conducted using MUSCLE. This data can be visualized here.
ClustalW2
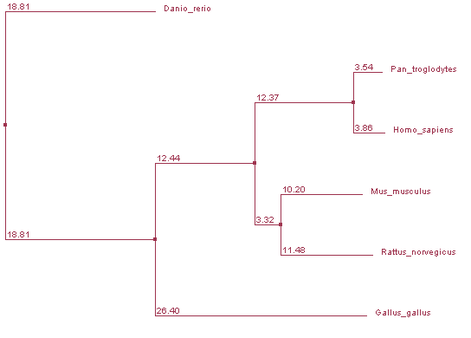
Phylogeny tree constructed by ClustalW2, using Neighbor Joining % Identity with distances.
Sequence alignment data (using ClustalW2) can be visualized here.
Parameters: Pairwise Alignment Options: Protein Weight Matrix: BLOSUM
Multiple Sequence Alignment Options: Protein Weight Matrix: BLOSUM
Sequence alignment data (using ClustalW2) can be visualized here.
Parameters: Pairwise Alignment Options: Protein Weight Matrix: BLOSUM
Multiple Sequence Alignment Options: Protein Weight Matrix: BLOSUM
T-Coffee
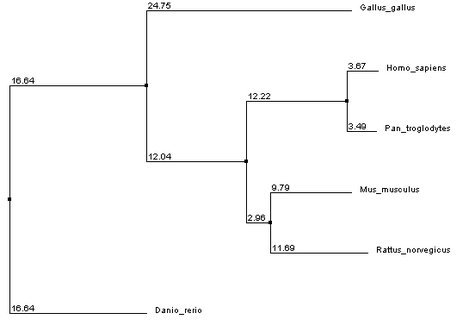
Phylogeny tree and sequence alignment data was conducted using T-Coffee. This data can be visualized here.
MUSCLE
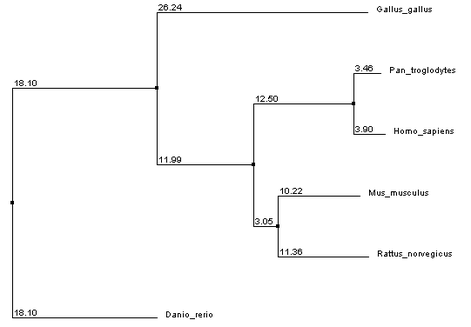
References
Site created by Valeri Lapacek
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
