_This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Microarray Technology
Biotechnology has advanced tremendously in the last decade. One tool used for genetic analysis is microarrays. Microarrays analyze the expression of DNA, that is, what genes are being expressed and in what amount. The proper amount of gene expression (amount of protein produced) is ideal for maintaining normal, healthy cellular functions. A microarray is a chip/slide that has numerous spots which contain DNA, cDNA, or oglionucleotides (short DNA strands). A chip may contain as many as 60,000 spots, thus analysis of 60,000 genes. Microarrays are useful because they can examine gene expression of many, many genes (spots) at once.
Gene Expression Omnibus (GEO) lists microarray experiments as well as the expression data. Microarray data can be analyzed in many ways. Many statistical approaches, like principle component analysis and hierarchical clustering, are used to measure patterns of similarity.
Gene Expression Omnibus (GEO) lists microarray experiments as well as the expression data. Microarray data can be analyzed in many ways. Many statistical approaches, like principle component analysis and hierarchical clustering, are used to measure patterns of similarity.
HLA-C Psoriasis Array
Figure 1
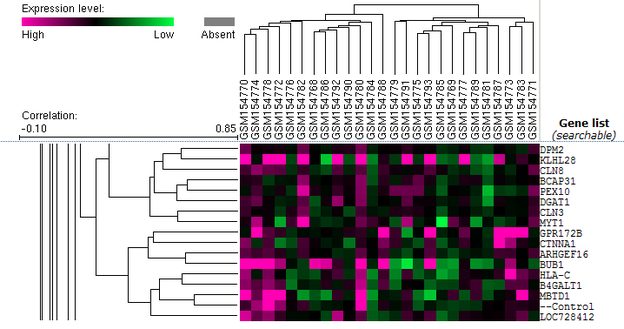
GEO identified an array analysis in order to study the molecular pathogenesis of psoriasis (see Figure 1). To visualize the GEO microarray data click here. The microarray data can be zoomed in (see Figure 2) for to see individual gene expression data. Clustering analysis often results in searching for functional similarities (like apoptosis) with the protein names in Princeton's GO Term Finder.
In this microarray experiment, HLA-C gene expression was examined in psoriasis patients via sampling psoriatic plaques and normal skin from 13 individuals. The study was searching for differential gene expression between psoriatic lesions and normal skin. The data was clustered (see black lines on left of images) and then analyzed by functions (using Princeton's GO Term Finder) to find differences in expression levels between normal and psoriatic skin. Click here to read published results.
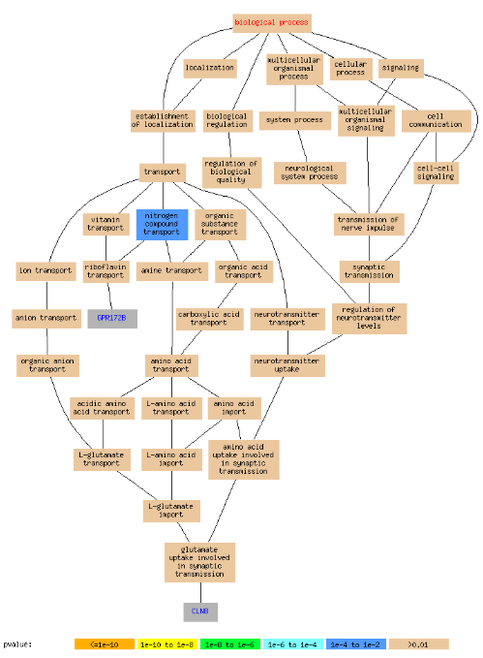
Figure 2 is clustered data including HLA-C. When surrounding proteins (see list of terms here) were plugged in Princeton's GO Term Finder a common function (nitrogen compound transport) was found ; however not involving HLA-C. The nitrogen compound transport function was found between two proteins: GPR172B and CLN8 (Figure 3). Figure 4 is a function tree summarizing how the GPR172B and CLN8 functions/paths are connected. This is a good example of how to used microarray data to find expression correlations between your gene of interest and other proteins.
In this microarray experiment, HLA-C gene expression was examined in psoriasis patients via sampling psoriatic plaques and normal skin from 13 individuals. The study was searching for differential gene expression between psoriatic lesions and normal skin. The data was clustered (see black lines on left of images) and then analyzed by functions (using Princeton's GO Term Finder) to find differences in expression levels between normal and psoriatic skin. Click here to read published results.
Figure 2 is clustered data including HLA-C. When surrounding proteins (see list of terms here) were plugged in Princeton's GO Term Finder a common function (nitrogen compound transport) was found ; however not involving HLA-C. The nitrogen compound transport function was found between two proteins: GPR172B and CLN8 (Figure 3). Figure 4 is a function tree summarizing how the GPR172B and CLN8 functions/paths are connected. This is a good example of how to used microarray data to find expression correlations between your gene of interest and other proteins.
References
1. Reischl J, Schwenke S, Beekman JM, Mrowietz U et al. Increased expression of Wnt5a in psoriatic plaques. J Invest Dermatol 2007 Jan;127(1):163-9.
2. GEO
3. Butte, A. The use and analysis of microarray data. Nature Reviews 2002 Dec;1:951-960.
4. Princeton's GO Term Finder
2. GEO
3. Butte, A. The use and analysis of microarray data. Nature Reviews 2002 Dec;1:951-960.
4. Princeton's GO Term Finder
_Site created by Valeri Lapacek
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012