This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is a domain?
Domains are units of a protein structure. A domain is made up of protein sequence (amino acids) and acts independent, in terms of function and structure, from the rest of the protein chain. Domains are independent in the sense that they fold independently from the rest of the amino acid chain and they can also function independently. A domain amino acid chain is first folded into alpha-helices and beta-sheets (secondary structure) and then compacted together (tertiary structure). It is the 3-D arrangement that creates functionality. Furthermore, domains can be genetically engineered so that they can combine and function with other proteins.
HLA-C Domains
Two protein domains were found using bioinformatic databases, Pfam and SMART. The two domains are Class I Histocompatibility antigen, domains alpha 1 and 2 (MHC_I) and Immunoglobulin C1-set (C1-set).
MHC_I
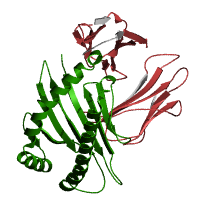
http://pfam.sanger.ac.uk/family/PF00129.13
Major Histocompatibility Complex (MHC) proteins are receptors located at cell surfaces. MHC proteins present antigens to T cells in order to create an immune response. Antigens are degraded protein fragments that are carried through the endoplasmic reticulum (E.R.) via TAP (transporter of antigenic peptides) proteins. MHC-I molecules bind the peptides from the E.R. and then become transported, via the Golgi apparatus. to cell surfaces (3,4).
There are two classes of MHC molecules, I and II. Class I molecules present intracellular antigenic peptides (~10 amino acids) to cytotoxic T cells, where as Class II molecules present exogenous antigenic peptides (~15 amino acids) to helper T cells.
There are two classes of MHC molecules, I and II. Class I molecules present intracellular antigenic peptides (~10 amino acids) to cytotoxic T cells, where as Class II molecules present exogenous antigenic peptides (~15 amino acids) to helper T cells.
C1-set
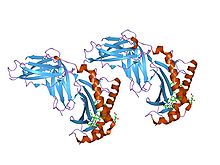
http://pfam.sanger.ac.uk/family/PF07654
MHC class I molecules have two chains: the heavy chain (MHC alpha) and the light chain (beta2-microglobulin). The alpha chain is the only chain that lies within the cell membrane. It consists of three extracellular domains, a transmembrane region, and a C-terminal cytoplasmic tail. The C1-set represents the C-terminal cytoplasmic tail domain. The C1-set domain are immunoglobulin-like domains, similar in structure and sequence. C1-set domains are most often involved in the immune system, resembling the antibody constant domain (5,6,7).
Pfam

http://pfam.sanger.ac.uk/search/sequence
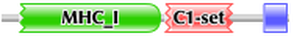
Pfam identified two domains, MHC_I and C1-set. MHC_I resulted in an e-value of 1e-98 with an alignment region of 26-203 (1). C1-set resulted in an e-value of 6.4e-21 with an alignment region of 218-294 (1). Figure shows two domains: MHC_I in green and C1-set in red. The blue box represents the region of the MHC I C family.
C1-set Pfam accession #: PF07654
MHC_I Pfam accession #: PF00129
C1-set Pfam accession #: PF07654
MHC_I Pfam accession #: PF00129
SMART
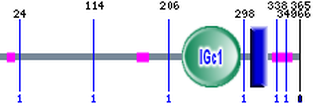
http://smart.embl-heidelberg.de/smart/show_motifs.pl
SMART identified two domains, IGc1 and transmembrane. IGc1 resulted in an e-value of 3.75e-26 with an alignment region of 222-293 (2). The transmembrane domain had no e-value, but had an alignment region of 305-327 (2). Figure shows two domains: IGc1 in green and a transmembrane domain in blue. The pink boxes are low complexity regions.
IGc1 SMART accession #: SM00407
IGc1 SMART accession #: SM00407
Analysis
Both, SMART and Pfam, domain analyses resulted in two domains. SMART identified the IGc1 and transmembrane domains, and Pfamidentified the MHC_I and C1-set domains. Without comparing the sequences, it is impossible to tell if the IGc1 domain is the same domain as MHC_I, and if the C1-set domain is the same as the transmembrane domain. Due to the different algorithms between the Pfam and SMART, the sequence alignment data is different.
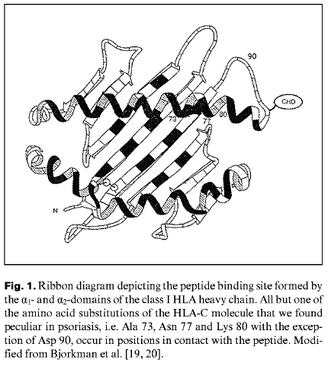
According to a paper by V. Brazzelli et al., there are 3 amino acid substitutions in the HLA-C (06) heavy chain region that predisposed psoriasis development. Two of these, Ala 73 and Asn 77, were located in the C pocket of the antigen binding groove and Lys 80 was in the F pocket (8). These pockets are significant in that they are contact positions (8). The location of the 3 mutations exist within one of the three extracellular domains of the C1-set.
According to a paper by V. Brazzelli et al., there are 3 amino acid substitutions in the HLA-C (06) heavy chain region that predisposed psoriasis development. Two of these, Ala 73 and Asn 77, were located in the C pocket of the antigen binding groove and Lys 80 was in the F pocket (8). These pockets are significant in that they are contact positions (8). The location of the 3 mutations exist within one of the three extracellular domains of the C1-set.
References
1. Pfam
2. SMART
3. Springer S, Doring K, Skipper JC, Townsend AR, Cerundolo V.
Fast association rates suggest a conformational change in the MHC class I molecule H-2Db upon peptide binding.
Biochemistry 37 3001-12 1998 [PubMed: 9485452]
http://dx.doi.org/10.1021/bi9717441
4. Paulsson KM. Evolutionary and functional perspectives of the major histocompatibility complex class I antigen-processing machinery. Cell. Mol. Life Sci. 61 2446-60 2004 [PubMed: 15526153]
http://dx.doi.org/10.1007/s00018-004-4113-0
5. Smith DK, Xue H.
Sequence profiles of immunoglobulin and immunoglobulin-like domains.
J. Mol. Biol. 274 530-45 1997 [PubMed: 9417933]
http://dx.doi.org/10.1006/jmbi.1997.1432
6. Pamer E, Cresswell P.
Mechanisms of MHC class I--restricted antigen processing.
Annu. Rev. Immunol. 16 323-58 1998 [PubMed: 9597133]
http://dx.doi.org/10.1146/annurev.immunol.16.1.323
7. Radosevich M, Ono SJ.
Novel mechanisms of class II major histocompatibility complex gene regulation.
Immunol. Res. 27 85-106 2003 [PubMed: 12637770]
http://dx.doi.org/10.1385/IR:27:1:85
8. V Brazzelli, et al. A pecucliar sequence motif in the aplpa-1-domain of the HLA-C molecule in psoriasis.
Dermatology. 200:2 pg. 99 (2000)
2. SMART
3. Springer S, Doring K, Skipper JC, Townsend AR, Cerundolo V.
Fast association rates suggest a conformational change in the MHC class I molecule H-2Db upon peptide binding.
Biochemistry 37 3001-12 1998 [PubMed: 9485452]
http://dx.doi.org/10.1021/bi9717441
4. Paulsson KM. Evolutionary and functional perspectives of the major histocompatibility complex class I antigen-processing machinery. Cell. Mol. Life Sci. 61 2446-60 2004 [PubMed: 15526153]
http://dx.doi.org/10.1007/s00018-004-4113-0
5. Smith DK, Xue H.
Sequence profiles of immunoglobulin and immunoglobulin-like domains.
J. Mol. Biol. 274 530-45 1997 [PubMed: 9417933]
http://dx.doi.org/10.1006/jmbi.1997.1432
6. Pamer E, Cresswell P.
Mechanisms of MHC class I--restricted antigen processing.
Annu. Rev. Immunol. 16 323-58 1998 [PubMed: 9597133]
http://dx.doi.org/10.1146/annurev.immunol.16.1.323
7. Radosevich M, Ono SJ.
Novel mechanisms of class II major histocompatibility complex gene regulation.
Immunol. Res. 27 85-106 2003 [PubMed: 12637770]
http://dx.doi.org/10.1385/IR:27:1:85
8. V Brazzelli, et al. A pecucliar sequence motif in the aplpa-1-domain of the HLA-C molecule in psoriasis.
Dermatology. 200:2 pg. 99 (2000)
Site created by Valeri Lapacek
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
Genetics 677 Assignment, Spring 2012
University of Wisconsin-Madison
Last Updated: 5/23/2012
